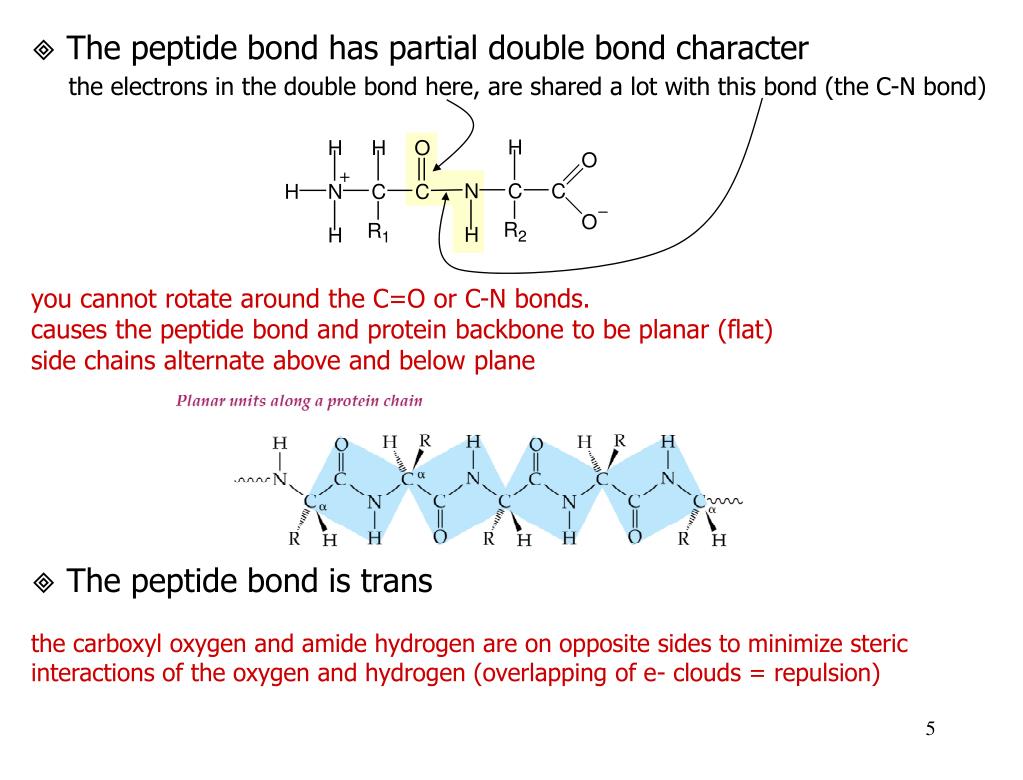
As any rotation is given by a rotation axis and degree of rotation, it can be plotted in 3D space as a vector along the rotation axis of length equal to the degree of rotation. The rotation is simply obtained as follows: First, rotate the entire protein, such that the donor peptide unit is aligned with the standard coordinate axes the subsequent rotation from this standard location of the donor to the acceptor is the 3D descriptor of the hydrogen bond. Here, we introduce a systematic 3D descriptor of main chain hydrogen bond geometry by assigning to each hydrogen bond between backbone C=O and N–H atoms a spatial rotation, which is evidently independent of the overall spatial orientation of the protein just as for dihedral angles. However, dihedral angles are not reliable when considering relative configurations, which are separated far apart along the backbone.Īs the geometric phase space of hydrogen bonds has large dimension a priori, 3D and 4D simplifications have captured only part of their geometry 5, 6. As hydrogen bonds are readily broken and reformed, they determine alternative conformations, and hence are also important for conformational changes of proteins.ĭihedral angles specify the backbone conformation of proteins by providing a complete 2D description of the progression from one peptide unit to the next along the backbone, as displayed in the well-known Ramachandran plots 4. 
Besides polypeptide assembly through covalent bonds, the structure is determined and stabilized by van der Waals interactions, hydrophobic packing, hydrogen bonds and ionic interactions. It is widely recognized that the function of a protein is intimately linked to the three-dimensional (3D) structure of its native folded state. Hydrogen bonds are of key importance in determining and fine-tuning molecular structure, interaction, function 2 and specificity of molecular recognition 3. It further provides a uniform vocabulary for comparison of protein structure near hydrogen bonds even between bonds in different proteins without alignment.Ī hydrogen bond 1 forms between an electronegative atom (the acceptor) and a hydrogen atom covalently bound to another electronegative atom (the donor). Strikingly, this rotational descriptor sampled over high-quality structures from the protein data base (PDB) concentrates into 30 localized clusters, some of which correlate to the common secondary structures and others to more special motifs, yet generally providing a unifying systematic classification of local structure around protein hydrogen bonds. 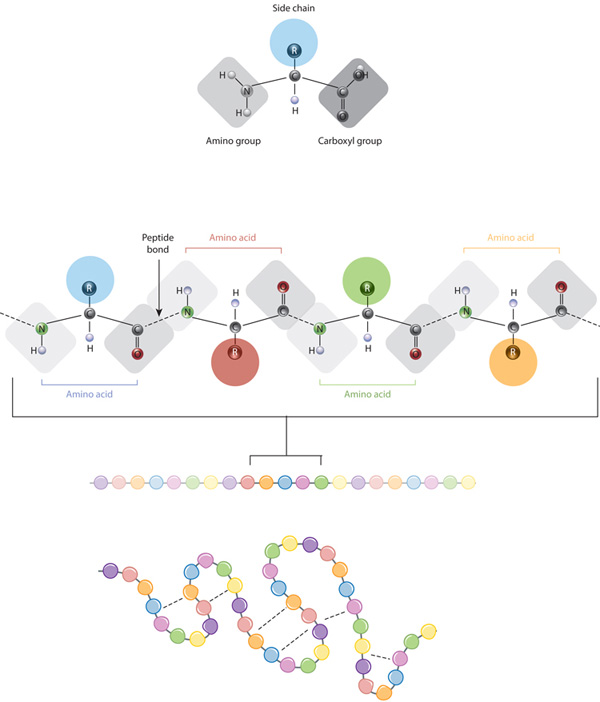
Here, we introduce the spatial rotation between hydrogen bonded peptide planes as a new descriptor for protein structure locally around a hydrogen bond. These, however, do not describe the conformations around hydrogen bonds, which can be non-local along the backbone and are of major importance for protein structure. The conformation of the backbone of each structure is locally at each C α effectively described by conformational angles resulting in Ramachandran plots. Proteins fold into three-dimensional structures, which determine their diverse functions.
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
